RNA-seq Analysis
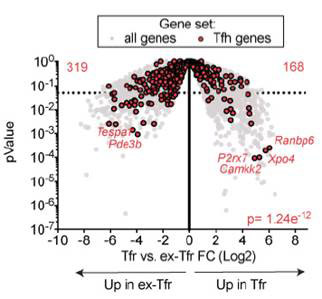
We provide a number of solutions for analyzing RNA-seq data, and we are here to help provide guidance throughout this process. There are two major solutions that we have available: The first option involves using to the schools' computing cluster (O2) this would involve using software such as the Tuxedo Suite (Link). For this option there are several ready built pipelines available for researchers to use. The second option involves using CLC Genomics workbench (Link) this is software package is installed on several of the machines in the IT suite (VSC 933). We have a robust workflow and tutorial along with experts to help you through this process.
Single Cell RNA-seq
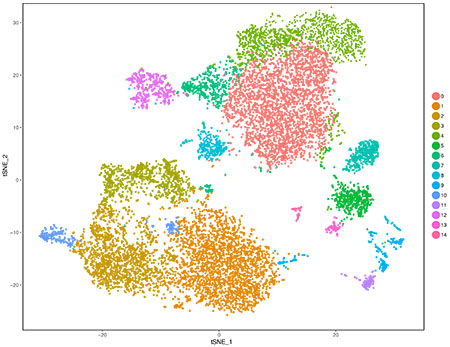
We provide advice and training support for both Cell Ranger and Seurat. In addition, we have connections to single cell cores in the Longwood Medical Area. This includes but is not limited to the Single Cell Genomics Core at BWH, as well as the Single Cell Core at HMS. We can help facilitate advice gathering from experts in the area.
Cell Ranger: We have a Cell Ranger pipeline for department members
Seurat: We provide general consulting help on how to use Seurat
Statistics
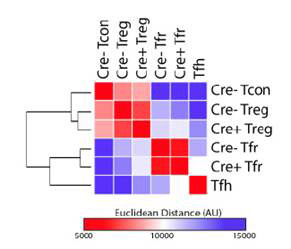
General statistics advice and consulting services are available for help with experimental design or how to compare results from expression analysis. We have experience with clinical trial experimental design, differential expression analysis, as well as general statistics.
R Programming
We provide guidance on how to configure and use R, the main workhorse of immunogenomics, and how to develop project-specific analysis paths with RStudio. In addition, we can follow the R introductory course with advice tailored to your project, in consultations that can range from brief visits to yearlong support.
Image Analysis
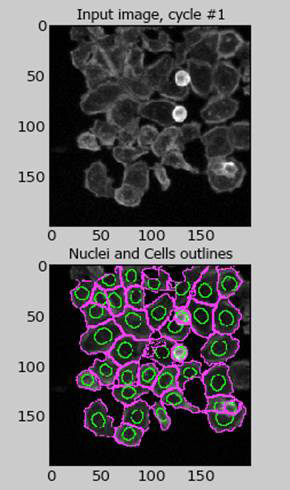
We have experience using Cell Profiler and Image analysis, as well as connections to many of the image analysis core facilities on campus.
Consulting
Not sure where to start? Just stop by VSC 933C. We can chat about your project and make recommendations to get you the help you need to proceed.